Why sequence a whole genome?
The main aim of sequencing the first human genome - back in 2005 - was to make a genomic map, indicating which genes are responsible for certain characteristics and hereditary diseases. But, in the end, this link is not quite so straightforward. Because you need to examine the genomes of thousands of people to be able to deduce anything useful. At the moment, this kind of work is done mainly in research centers, where the sequencing of whole genomes is used as an important research tool.
But reading full genomes (of human cells, viruses and bacteria) is also of value for establishing diagnoses, as well as for selecting and monitoring treatments. Experts believe that DNA-sequencing can become one of the most important tools used by doctors and specialists in their work – just as a standard blood analysis is now. “You can start by looking for hereditary diseases – which is already done today for breast cancer (BRCA 1 and 2 genes) – but sequencing the genome is also of interest for other, non-hereditary cancers,” explains Liesbet Lagae, program director life science technologies at imec. “Trouble is, a tumor is often made up of different sorts of cells, which have each undergone different mutations. At the moment, we can only look at the ‘average’ genome of the tumor at one specific time. This is because ‘deep sequencing’ is still too expensive and time-consuming. But if we were able to sequence the genome of a cell, quickly and cheaply, we could monitor the various types of cancer cells very precisely over a specific period of time and so treat them better.”
“This sort of single-cell sequencing is important for deciding on the correct cancer treatment, such as with multiple myeloma,” adds Wilfried Verachtert, director ExaScience Life Lab. “With some cancers, doctors can see that the cell types in the tumor mutate during treatment. And if that happens, a new kind of treatment has to be started. But if we were able to screen the composition of the various tumor cells with greater frequency, it would make it much easier to treat these more complex cancers.”
Our blood not only contains our own hereditary material, but also that of viruses and bacteria. Detecting these genomes is of value for detecting and treating infectious diseases, such as Ebola or new strains of influenza.
Despite the enormous possibilities opened up by DNA-sequencing, it is still not used routinely in day-to-day hospital practice,. In fact there are very few FDA-approved diagnostic tests based on DNA-sequencing. Why is that? Because the process is still too expensive and cumbersome.

"At the moment, we can only look at the ‘average’ genome of the tumor at one specific time. This is because ‘deep sequencing’ is still too expensive and time-consuming. But if we were able to sequence the genome of a cell, quickly and cheaply, we could monitor the various types of cancer cells very precisely over a specific period of time and so treat them better.”
The $1000 genome: an unprecedented success
In 2005, the ‘National Human Genome Research Institute’ (NHGRI) in America launched the $1000 genome project. At that time, the Human Genome Project – through which the first human genome had been examined for the first time – had just been completed. It had cost $3 billion and hundreds of researchers had spent 13 years working on it. So now it was time for the next milestone: reducing those $3 billion to just $1000 per genome. An impossible task – or so it seemed at the time.
More than 100 research groups from both the academic and business worlds submitted projects to the NHGRI and received grants to make the $1000 genome a reality. “The project was a great success and, today, sequencing a full genome does indeed cost $1000. And in the years ahead, the cost will fall even further, maybe to $100 and even $10,” says Liesbet Lagae. “Imec is also involved in this progress. At the moment we are working in conjunction with more than five major manufacturers of sequencing tools to make their sequencing chip faster, more efficient and cheaper. And to do that, we are using our expertise in photonics, sensors, integration and chip production.”
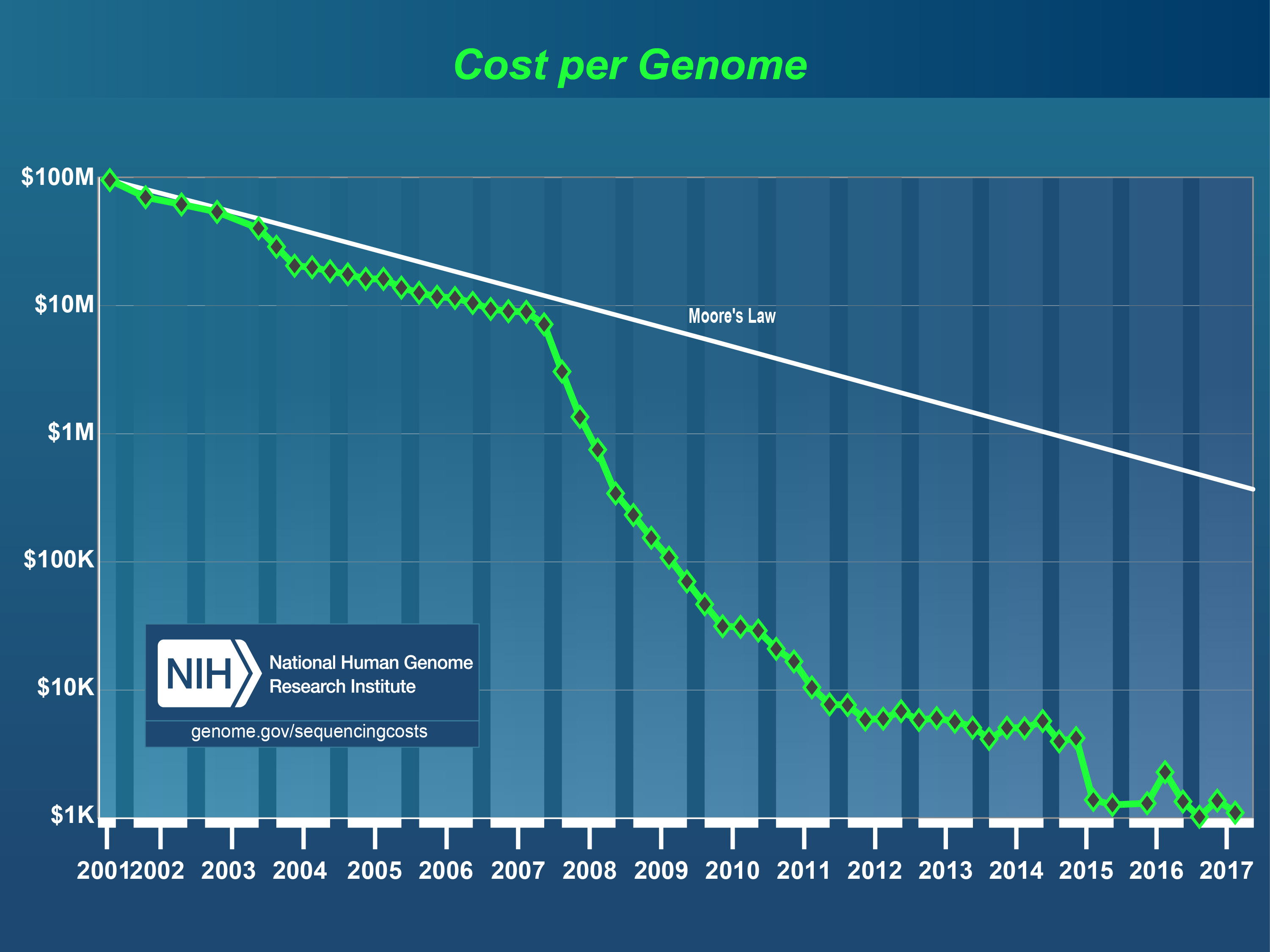
The cost per genome, as measured by the NHGRI. The way the cost is progressing can be compared with the hypothetical cost in the way that technology follows the predictions of Moore’s Law. Moore’s Law is used in the chip industry as the long-term trend by which it is assumed that the calculating power of a computer can double every two years. The chip industry is doing everything it can to maintain this trend and in doing so has been responsible for many revolutionary developments. The technology for DNA-sequencing appears to be capable of advancing even faster than Moore’s Law.
Imec's roadmap towards the 10-dollar genome.
But all of these sequencing tools – that are able to read the genome for $1000 – are not the whole picture. The DNA sample placed into these tools first of all has to be created from the patient’s blood by specially trained operators in a separate laboratory. And the raw data produced by the sequencing tools cannot simply be used without further analysis to make a diagnosis or prescribe a treatment. This means that before GPs and specialists can use DNA-sequencing as a standard tool, these two additional steps also need to be tackled.
Changes over time of the contribution of the various steps in sequencing to the cost of sequencing a genome (Sboner et al. Genome Biology 2011, 12:125).
Can we prepare the sample in a small lab-on-chip?
At the moment, the sample to be used in sequencing tools has to be prepared in a specialized lab. But suppose we could replace that laboratory with a lab-on-chip? “Preparing the sample currently costs around $200,” says Liesbet Lagae. “Previously, when reading the DNA still cost a few thousand dollars, this was a negligible cost. But as more and more progress is made and the cost of the sequencing itself continues to fall, it is now time to tackle the cost of preparation. This can be achieved by designing a microfluidics chip that carries out these various steps – as they would be done in the laboratory – but much faster, in a more compact way and using fewer reagents (which would also make it cheaper).”
Imec aims to use a silicon platform to do this.
“We have already developed the various building blocks needed for this chip: PCR, DNA extraction, mixing reagents, synthetizing the DNA and so on. And we can combine these in a complex flow, as we have already done for Panasonic and their lab-on-chip for SNP detection. This means that our microfluidics platform, based on silicon, is definitely able to take DNA-sequencing that little step further.”
And yet it is still not easy. “The big challenge is that the diagnosis has to be very precise and accurate,” adds Liesbet Lagae. “The flow for the sample preparation cannot fail under any circumstances and none of the building blocks can go wrong either. In case of the sequencing step, accuracy is built into the process by repeating the sequencing of the same DNA fragments many, many times (up to as many as 60 times!). However, any inaccuracy in the sample preparation is much more difficult to compensate.
Will we be able to produce a comprehensible dashboard for the doctor in just a matter of hours?
We can already read a genome in just one day, but typically it takes another two weeks to convert the data into useful information for researchers and doctors. Which is too long if we want to use DNA-sequencing as a diagnostic tool. With many tests you only have to analyze a small part of the genome to be able to start drawing very simple conclusions. And it is precisely these applications that are now being translated into clinical practice in the form of FDA-approved diagnostic tests. Yet by doing this we are only analyzing the tip of the iceberg.
“In a previous project, with Janssen Pharmaceutica, Intel and UGent, we used our software expertise in high-performance computing and machine learning to substantially speed up what is known as the ‘alignment step’ that takes place after sequencing (ed.: all of the DNA fragments are then put together to create the full genetic map of our 23 pairs of chromosomes),” explains Wilfried Verachtert. “And thanks to the open-source software that was developed as part of the project – elPrep and Halvade – this step no longer takes 5 days, but can be done in just 1½ hours.”
“Now what we need to do is also speed up the ‘variant calling’ step and make it usable for doctors,” he adds. “In this process, the computer goes looking for the differences between a specific genome and the genomes of other patients. Our aim is to use machine learning to detect patterns in the data from thousands of patients who have a specific disease. That way the computer is able to ‘learn’ to identify the illness in new samples.
The ultimate goal is to produce a computer chip that tells the doctor what relevant information there is to be found in the sequencing data in a form that the doctor can interpret quickly to make the correct diagnosis or to decide on what the best treatment might be.”
And while we’re still dreaming, would it be possible to expand the point-of-care sequencing devices already on the market (such as MiniSeq from Illumina or the Minion from Oxford Nanopore) to include a lab-on-chip for preparing samples, plus an artificical-intelligence-chip for interpreting the data? If we could, a doctor would then simply use this desktop tool to create a personalized treatment for the patient using just one drop of blood to read his or her DNA. In any event, imec will be doing everything it can, in conjunction with interested companies, to make this dream become a reality.
Want to know more?
- Read more about how imec and Panasonic are working together on the SNP chip
- This press release explains how the elPrep software speeds up genome analysis
- More information about the Exascience Life Lab
- Last month’s magazine featured everything you need to know about machine learning and artificial intelligence:
- Would you like to work with imec on microfluidic chips or machine learning? If you would, then let us know via this contact form.

Liesbet Lagae is co-founder and currently Program Director of the Life Science Technologies in imec. In this role, she oversees the emerging R&D, the public funded activities and early business creation. She holds a PhD degree from the KU Leuven, Belgium for her work on Magnetic Random Access Memories obtained under an IWT grant.
As a young group leader, she has initiated the field of molecular and cellular biochips leveraging silicon technologies at imec, Belgium. The life science program has grown from emerging activities to a mature business line that provides smart silicon chip solutions to the life science industry. Applications include medical diagnostics, point-of-care solutions, DNA sequencing, cytometry, bioreactors, neuroprobes, implants.

Wilfried Verachtert graduated as Lic. Informatics at the Brussels Free University. He continued his career as researcher at the Brussels Free University on Parallel Programming Languages. He did join a spin-off software company “MediaGenix” as partner & CTO. 14 years ago he joined imec as Group Director Digital Components. 7 years ago, he co-launched - and became the director of - the Exascience Life Lab, a collaboration between Imec, Intel, Janssen Pharmaceutic and the 6 Flemish universities to do applied research on High Performance Computing for Life Sciences. Since May this year, became PMTS (Principal Member of the Technical Staff) Data & AI at Imec to work on the longer term vision of Data and AI.
Published on:
1 October 2018














